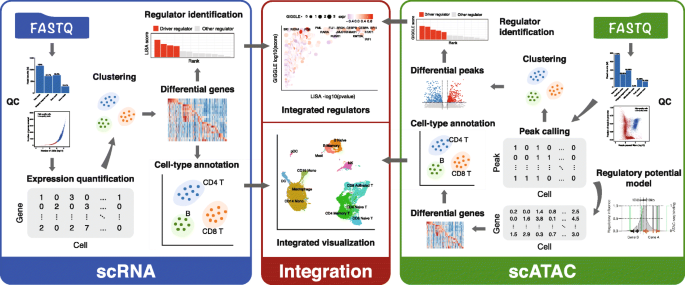
Integrative analyses of single-cell transcriptome and regulome using MAESTRO | Genome Biology | Full Text

Integrative analysis of single-cell genomics data by coupled nonnegative matrix factorizations | PNAS

Single-Cell RNA Sequencing and Assay for Transposase-Accessible Chromatin Using Sequencing Reveals Cellular and Molecular Dynamics of Aortic Aging in Mice | Arteriosclerosis, Thrombosis, and Vascular Biology
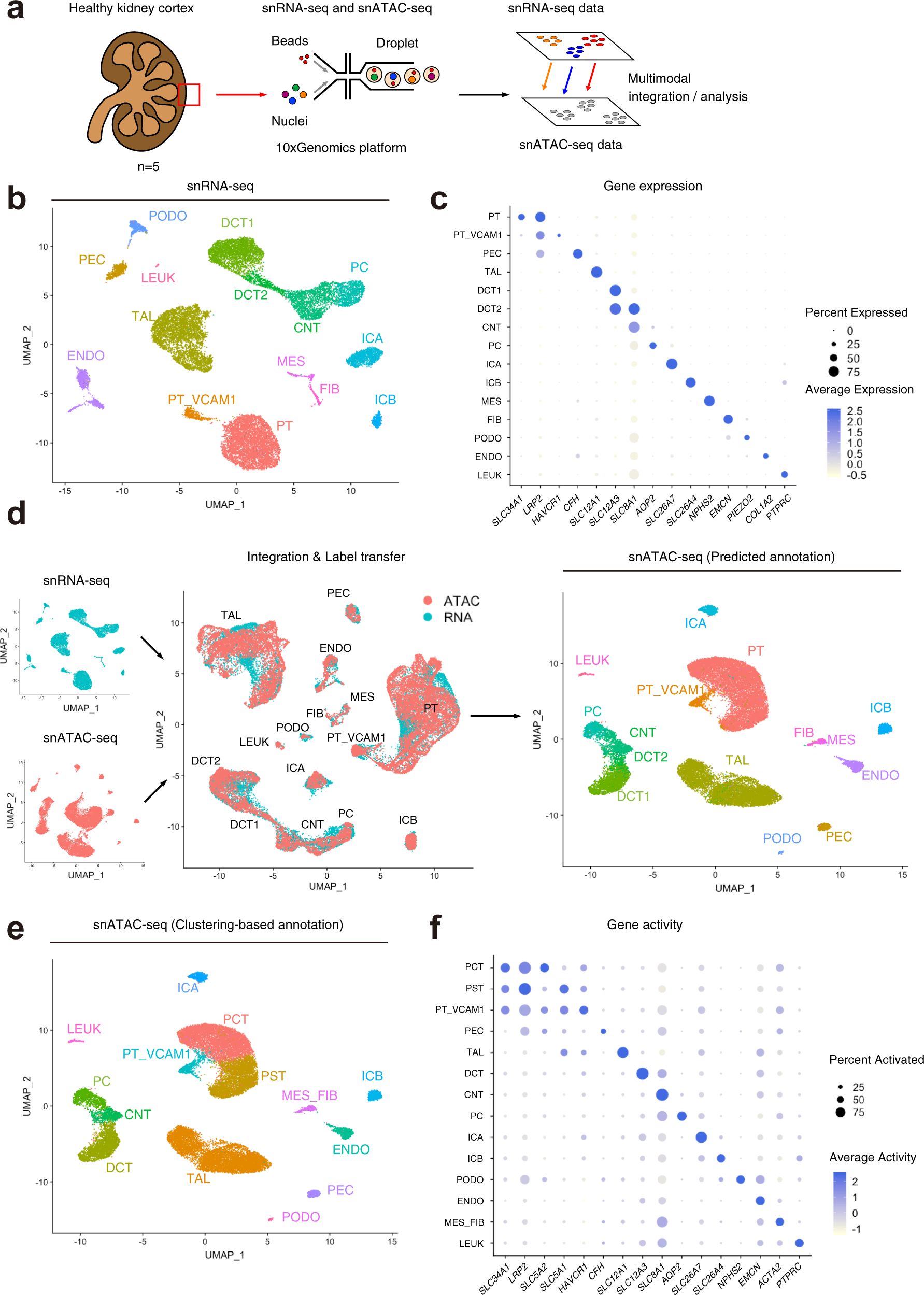
Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney | Nature Communications
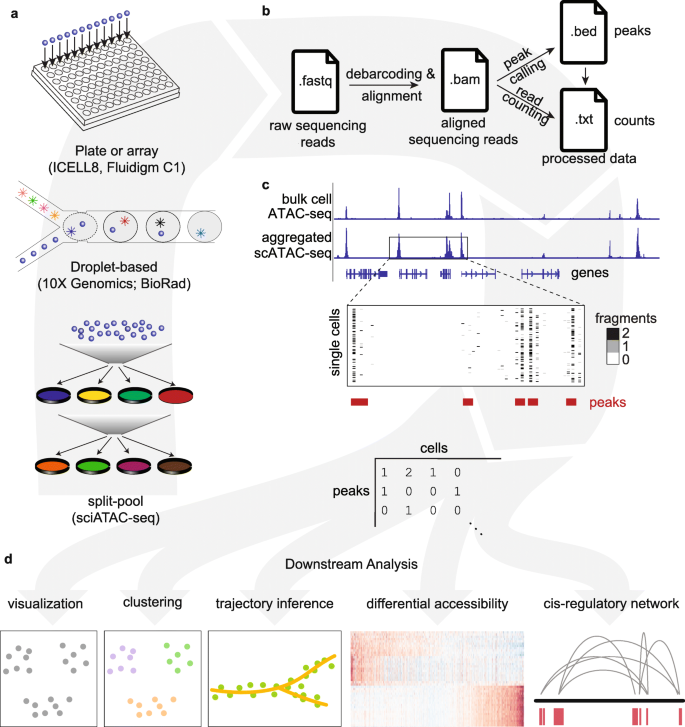
Assessment of computational methods for the analysis of single-cell ATAC-seq data | Genome Biology | Full Text

Simultaneous trimodal single-cell measurement of transcripts, epitopes, and chromatin accessibility using TEA-seq | eLife

BGI Genomics on Twitter: "Novel single cell research based on BGI's DNBseq™ technology published in @NatureComms. scCAT-seq integrates single-cell ATAC- seq & RNA-seq to provide an accurate genome-wide measure of chromatin accessibility &

Identification of genomic enhancers through spatial integration of single‐ cell transcriptomics and epigenomics | Molecular Systems Biology
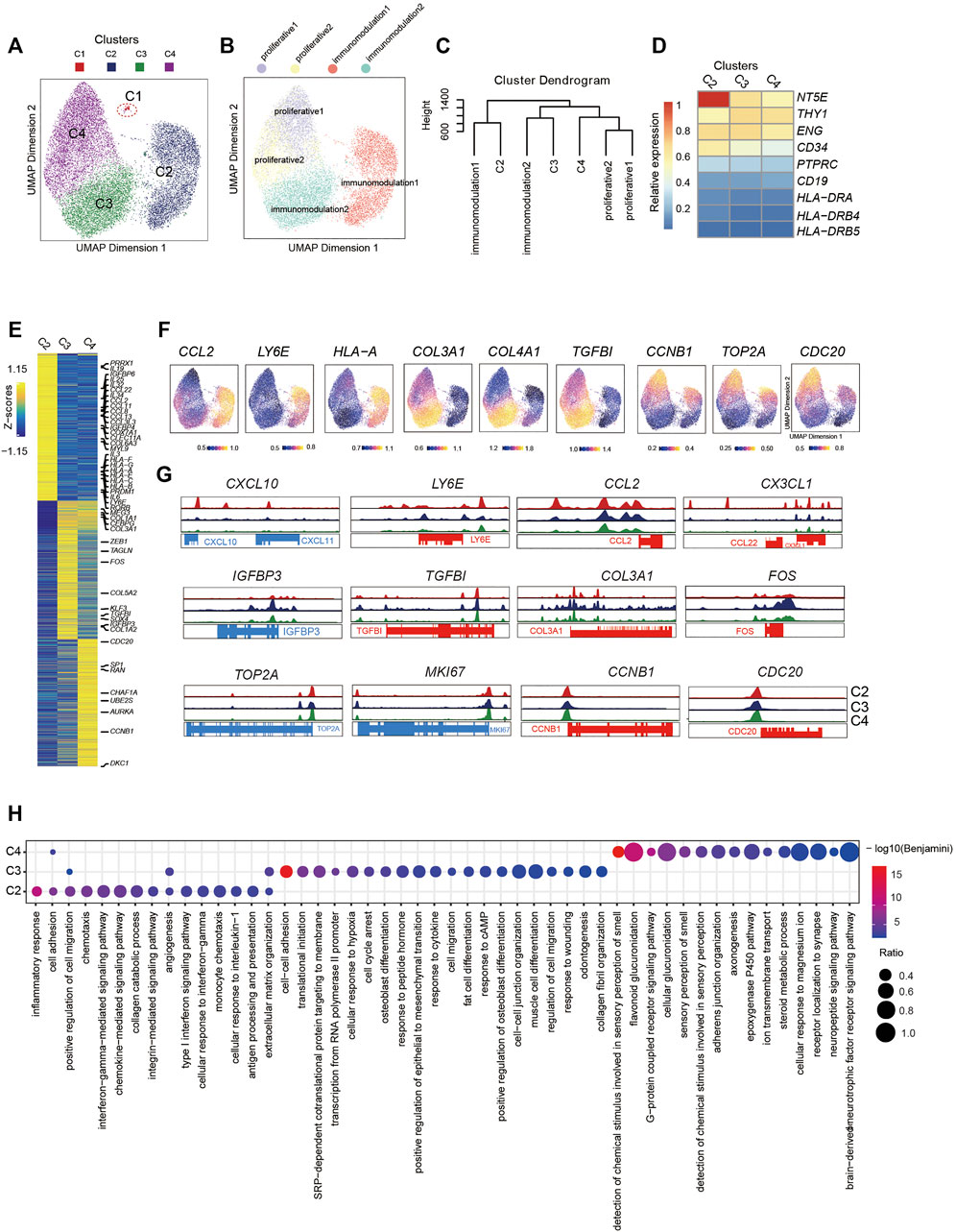
Frontiers | Integrative Single-Cell RNA-Seq and ATAC-Seq Analysis of Mesenchymal Stem/Stromal Cells Derived from Human Placenta
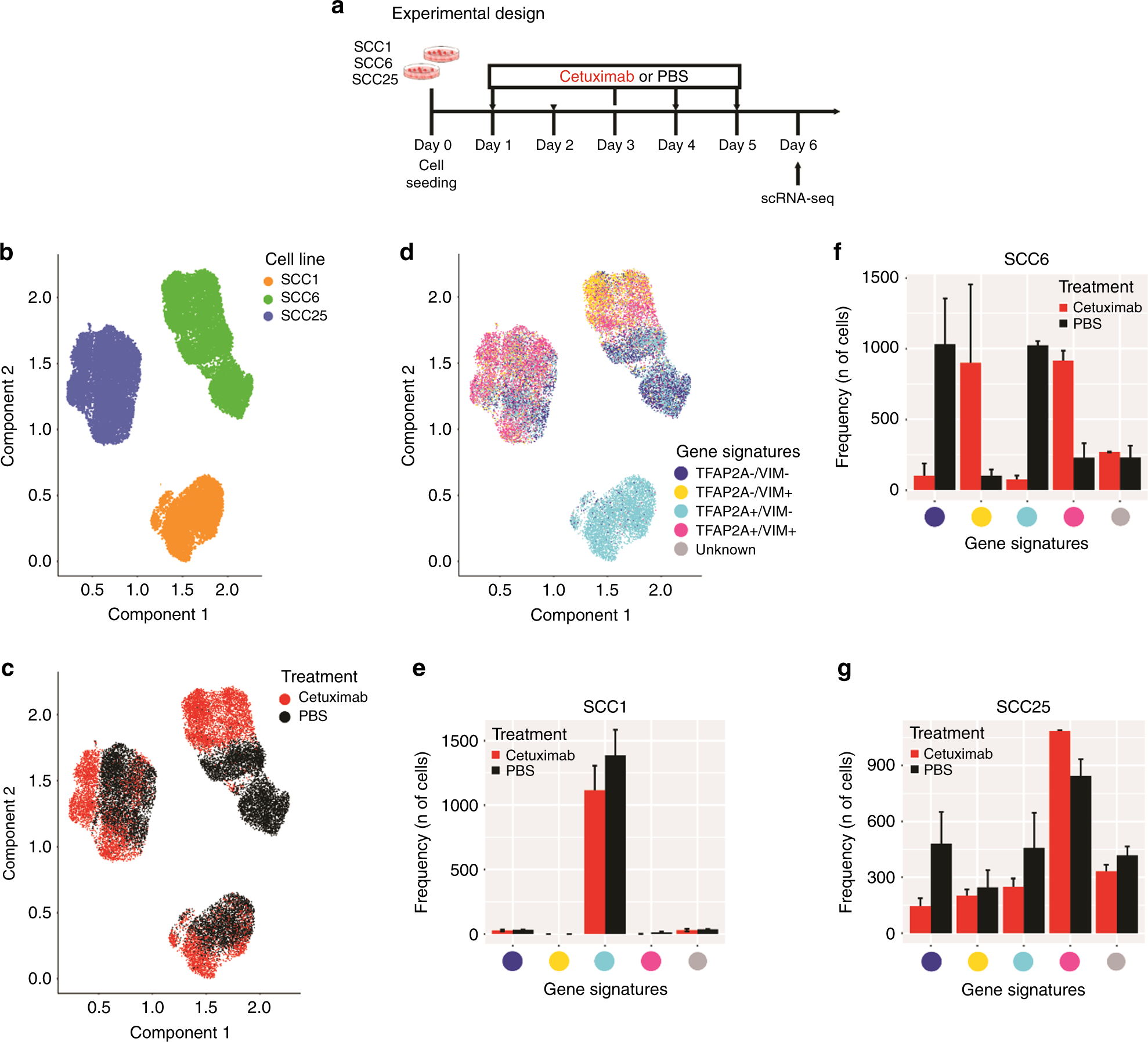
Integrated single-cell and bulk gene expression and ATAC-seq reveals heterogeneity and early changes in pathways associated with resistance to cetuximab in HNSCC-sensitive cell lines | British Journal of Cancer

Identification of genomic enhancers through spatial integration of single‐ cell transcriptomics and epigenomics | Molecular Systems Biology
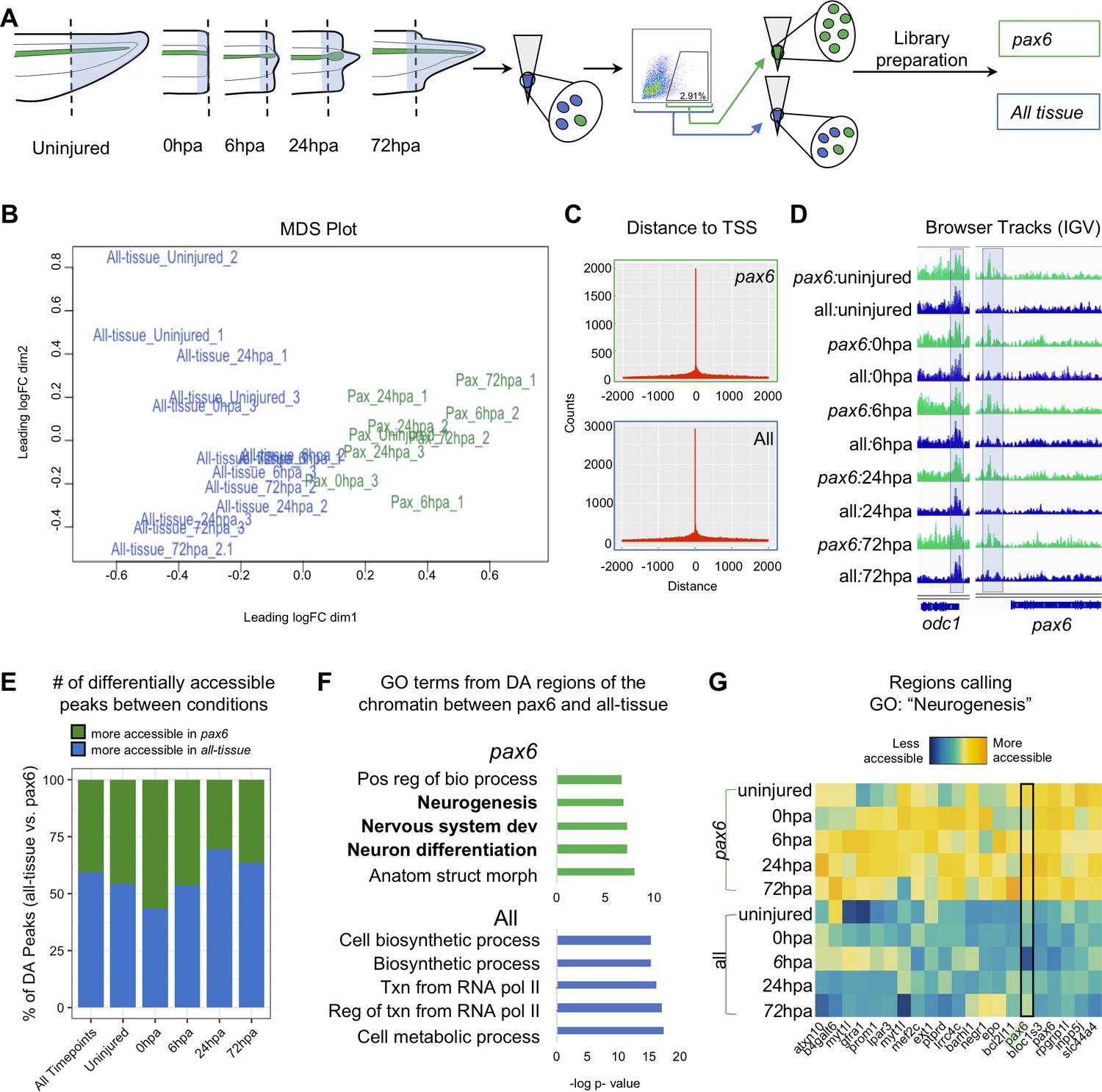
Chromatin accessibility dynamics and single cell RNA-Seq reveal new regulators of regeneration in neural progenitors | eLife

Joint profiling of chromatin accessibility and gene expression in thousands of single cells | Science

Single-cell ATAC sequencing analysis: From data preprocessing to hypothesis generation - ScienceDirect

Single-Cell Epigenomics and Functional Fine-Mapping of Atherosclerosis GWAS Loci | Circulation Research

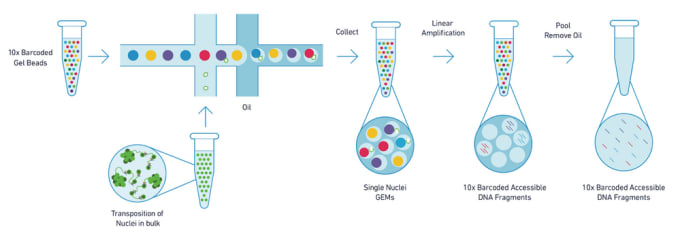
Single-cell ATAC-seq. (a) The workflow of scATAC-seq to measure single... | Download Scientific Diagram
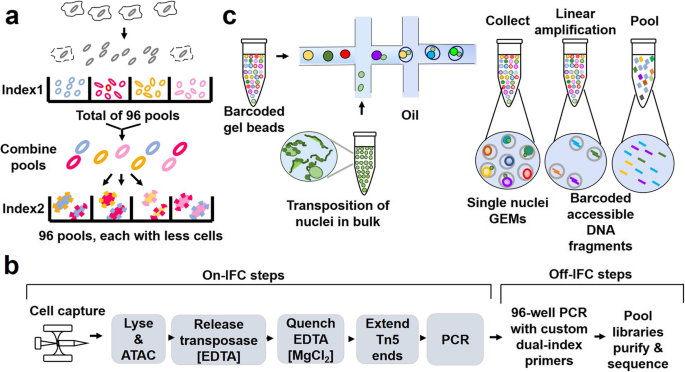
Profiling chromatin regulatory landscape: insights into the development of ChIP-seq and ATAC-seq | SpringerLink